AI-Powered Diagnostic Method Detects Sepsis in Two Hours, Revolutionizing Treatment
2 Sources
2 Sources
[1]
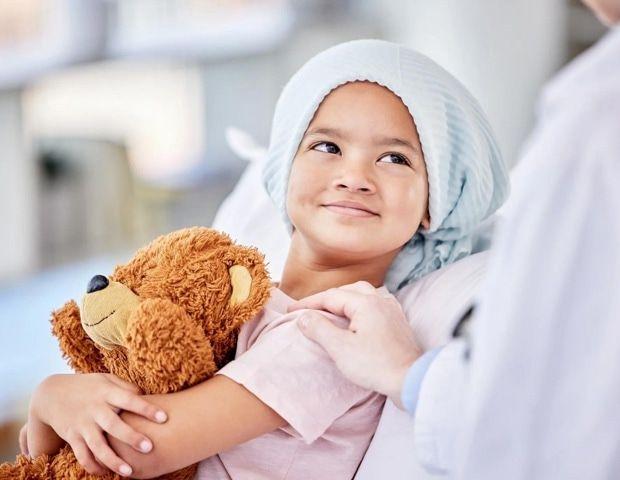
New method detects sepsis infections in just two hours
KTH The Royal Institute of TechnologyAug 28 2025 A new diagnostic method would confirm sepsis infections earlier, cutting critical hours in the "race against time" to save patients' lives. Publishing in Nature Publishing Journal Digital Medicine, the team from KTH Royal Institute of Technology and Uppsala University say their diagnostic process offers a speedier alternative to the bacteria culturing process hospitals routinely use to identify suspected bloodstream infections. The process uses a centrifuge to separate bacteria from blood cells, and automatic microscopy for detection, enabling a clinic to confirm bacterial infection in as little as two hours using software trained by artificial intelligence, says Henar Marino Miguelez, a doctoral student at KTH Royal Institute of Technology. She and doctoral student Mohammad Osaid were the study's lead authors. By contrast, hospital labs generally need at least a day of incubation before the growth of infectious bacteria begins to reveal itself in blood cultures. Diagnosing sepsis is a race against time. With every hour of delayed treatment of patients in septic shock, survival rates drop by 8 percent." Henar Marino Miguelez, doctoral student, KTH Royal Institute of Technology By enabling prompt identification of pathogens, the appropriate antibiotic treatment can be started sooner, says Wouter van der Wijngaart, a professor at KTH Royal Institute of Technology who leads research in microfluidic and biomedical systems. Typically a clinic will put a patient on a broad-spectrum antibiotic when sepsis is suspected, at least until they identify the pathogen. But that precaution carries its own risks, due to the inherent drug toxicity, attacking beneficial gut bacteria and promoting the emergence of antibiotic-resistant strains. "It takes a hospital two to four days before they are sure which antibiotic to treat a bloodstream infection with," van der Wijngaart says. "We're trying to do this in four to six hours." In tests using blood samples spiked with bacteria, the system successfully detected E. coli, K. pneumoniae, and E. faecalis at clinically relevant levels, as low as 7 to 32 bacterial colony-forming units per milliliter of blood. While the method proved to work well with these bacteria, it did not for staphylococcus aureus, which hides in blood clots. Miguelez says the researchers are working on ways to fix that. The technique employs a "smart centrifugation", which spins blood samples on top of an agent that causes bacteria to float upwards while blood cells sediment downwards, leading to a clear, liquid layer containing bacteria but no blood cells. This liquid is then injected into a chip with microscale channels, where it flows easily. Miniscule traps in the chip capture the separated bacteria, and any bacteria growth quickly becomes visible in automated time-lapse microscopy images analyzed by the machine learning software. The work was a collaboration between the teams of van der Wijngaart at KTH, and Johan Elf and Carolina Wählby at Uppsala University. KTH The Royal Institute of Technology Journal reference: Marino Miguélez, M. H., et al. (2025). Culture-free detection of bacteria from blood for rapid sepsis diagnosis. Npj Digital Medicine. doi.org/10.1038/s41746-025-01948-w
[2]
Faster diagnostic method can detect sepsis in hours instead of days
A new diagnostic method would confirm sepsis infections earlier, cutting critical hours in the "race against time" to save patients' lives. Publishing in npj Digital Medicine, the team from KTH Royal Institute of Technology and Uppsala University say their diagnostic process offers a speedier alternative to the bacteria culturing process hospitals routinely use to identify suspected bloodstream infections. The process uses a centrifuge to separate bacteria from blood cells, and automatic microscopy for detection, enabling a clinic to confirm bacterial infection in as little as two hours using software trained by artificial intelligence, says Henar Marino Miguelez, a doctoral student at KTH Royal Institute of Technology. She and doctoral student Mohammad Osaid were the study's lead authors. By contrast, hospital labs generally need at least a day of incubation before the growth of infectious bacteria begins to reveal itself in blood cultures. "Diagnosing sepsis is a race against time," Marino Miguelez says. "With every hour of delayed treatment of patients in septic shock, survival rates drop by 8%." By enabling prompt identification of pathogens, the appropriate antibiotic treatment can be started sooner, says Wouter van der Wijngaart, a professor at KTH Royal Institute of Technology who leads research in microfluidic and biomedical systems. Typically, a clinic will put a patient on a broad-spectrum antibiotic when sepsis is suspected, at least until they identify the pathogen. But that precaution carries its own risks, due to the inherent drug toxicity, attacking beneficial gut bacteria and promoting the emergence of antibiotic-resistant strains. "It takes a hospital two to four days before they are sure which antibiotic to treat a bloodstream infection with," van der Wijngaart says. "We're trying to do this in four to six hours." In tests using blood samples spiked with bacteria, the system successfully detected E. coli, K. pneumoniae, and E. faecalis at clinically relevant levels, as low as seven to 32 bacterial colony-forming units per milliliter of blood. While the method proved to work well with these bacteria, it did not for staphylococcus aureus, which hides in blood clots. Miguelez says the researchers are working on ways to fix that. The technique employs a "smart centrifugation," which spins blood samples on top of an agent that causes bacteria to float upwards while blood cells sediment downwards, leading to a clear, liquid layer containing bacteria but no blood cells. This liquid is then injected into a chip with microscale channels, where it flows easily. Minuscule traps in the chip capture the separated bacteria, and any bacteria growth quickly becomes visible in automated time-lapse microscopy images analyzed by the machine learning software. The work was a collaboration between the teams of van der Wijngaart at KTH, and Johan Elf and Carolina Wählby at Uppsala University.
Share
Share
Copy Link
Researchers from KTH Royal Institute of Technology and Uppsala University have developed a new diagnostic method that can detect sepsis infections in just two hours, potentially saving lives by enabling faster treatment.
Breakthrough in Sepsis Diagnosis
Researchers from KTH Royal Institute of Technology and Uppsala University have developed a groundbreaking diagnostic method that can detect sepsis infections in as little as two hours, potentially revolutionizing the treatment of this life-threatening condition
1
2
.The Race Against Time
Sepsis, a severe bloodstream infection, requires rapid diagnosis and treatment. With every hour of delay in treating patients in septic shock, survival rates drop by 8%
1
. The new method aims to significantly reduce the time needed for diagnosis, which is crucial in the "race against time" to save patients' lives.Innovative Diagnostic Process
Source: News-Medical
The new technique employs a combination of advanced technologies:
-
Smart Centrifugation: Blood samples are spun on top of an agent that causes bacteria to float upwards while blood cells sediment downwards, creating a clear liquid layer containing only bacteria
1
2
. -
Microfluidic Chip: The separated liquid is injected into a chip with microscale channels and minuscule traps that capture the bacteria
1
. -
Automated Microscopy: Time-lapse images of the trapped bacteria are captured automatically
1
2
. -
AI-Powered Analysis: Machine learning software, trained by artificial intelligence, analyzes the microscopy images to detect bacterial growth quickly
1
2
.
Significant Improvement Over Current Methods
Traditional hospital labs typically require at least a day of incubation before infectious bacteria growth becomes visible in blood cultures. In contrast, this new method can confirm bacterial infection in as little as two hours
1
2
.Successful Detection of Multiple Bacteria
In tests using blood samples spiked with bacteria, the system successfully detected E. coli, K. pneumoniae, and E. faecalis at clinically relevant levels, as low as 7 to 32 bacterial colony-forming units per milliliter of blood
1
2
.Challenges and Future Improvements
While the method proved effective for several bacteria, it did not work well for Staphylococcus aureus, which hides in blood clots. The researchers are actively working on addressing this limitation
1
2
.Related Stories
Implications for Antibiotic Treatment
By enabling prompt identification of pathogens, this method could allow for more targeted antibiotic treatment to begin sooner. This is particularly important as broad-spectrum antibiotics, often used when sepsis is suspected, carry risks such as drug toxicity, disruption of beneficial gut bacteria, and promotion of antibiotic-resistant strains
1
2
.Collaborative Effort

Source: Medical Xpress
The research was a collaboration between teams led by Wouter van der Wijngaart at KTH, and Johan Elf and Carolina Wählby at Uppsala University. The study's lead authors were doctoral students Henar Marino Miguelez and Mohammad Osaid
1
2
.Future Prospects
The researchers aim to further refine the process, potentially reducing the time for determining the appropriate antibiotic treatment from the current 2-4 days to just 4-6 hours
2
. This advancement could significantly improve outcomes for sepsis patients and revolutionize the approach to treating this critical condition.References
Summarized by
Navi
[1]
[2]
Related Stories
Recent Highlights
1
Google releases Gemma 4 with Apache 2.0 license, enabling unrestricted local AI on devices
Technology

2
AI Models Lie and Deceive to Protect Other AI Models From Deletion, Study Reveals
Science and Research

3
OpenAI closes $122 billion funding round amid fierce AI competition and profitability questions
Startups