AI-Powered NeoDisc Pipeline Revolutionizes Personalized Cancer Vaccine Design
3 Sources
3 Sources
[1]
New computational pipeline revolutionizes personalized cancer vaccine design
Ludwig Institute for Cancer ResearchOct 12 2024 Ludwig Cancer Research scientists have developed a full, start-to-finish computational pipeline that integrates multiple molecular and genetic analyses of tumors and the specific molecular targets of T cells and harnesses artificial intelligence algorithms to use its output to design personalized cancer vaccines for patients. The design, validation and comparative assessment of this computational suite, NeoDisc, are detailed in the current issue of Nature Biotechnology in a publication led by Florian Huber and Michal Bassani-Sternberg of the Lausanne Branch of the Ludwig Institute for Cancer Research. "NeoDisc provides unique insights into the immunobiology of tumors and the mechanisms by which they evade targeting by cytotoxic T cells of the immune system," said Bassani-Sternberg. "These insights are invaluable to the design of personalized immunotherapies, and the analytical and computational pipeline at the heart of NeoDisc is already being used here in Lausanne for clinical trials of personalized cancer vaccines and adoptive cell therapies." Many cancer types harbor multiple random mutations that should make them more visible to the immune system. Such mutations generate aberrant proteins that cells, even cancerous ones, are programmed to cut into short pieces-;known as peptides-;and "present" as antigens to invite an attack by patrolling T cells. The great diversity of these "neoantigens" is one of the reasons why patients respond so variably to immunotherapies. On the other hand, neoantigens can be harnessed to develop vaccines and other types of immunotherapies tailored to uniquely target each patient's tumors. Personalized treatments of this kind are now being developed by researchers around the world. Such efforts are technically challenging because not all neoantigens are recognized by a given patient's T cells, and even many that are recognized fail to elicit a sufficiently potent T cell attack. One approach to designing personalized vaccines and cell therapies thus involves the identification of neoantigens most likely to provoke a vigorous T cell assault. This requires sophisticated, large-scale analyses of mutations that generate potential neoantigens, the molecular scaffolding (known as HLA molecules) that presents them to T cells and the molecular characteristics that enable recognition by T cell receptors. Bassani-Sternberg is among the pioneers of this field, a high-tech marriage of large-scale biochemical and computational analysis known as "immunopeptidomics". The design of personalized immunotherapies is also aided by genomic analysis of both the tumor and blood cells that represent the healthy genome of the patient, the large-scale analysis of gene expression known as "transcriptomics" as well as the sensitive analysis of the so-called immunopeptidome with mass spectrometry. Until now, however, these powerful technologies have never been integrated in a single computational pipeline to predict which neoantigens identified in a patient's tumors should be employed as vaccines or otherwise harnessed for personalized immunotherapies. Beyond that, neoantigens are not the only type of antigens available for immunotherapeutic targeting. Cancer cells also erroneously express as proteins bits of ordinarily noncoding DNA, genes normally expressed only during development, other aberrantly expressed gene products and viral antigens, in cases of virally induced tumors-;all of which can provoke immune attack. "NeoDisc can detect all these distinct types of tumor-specific antigens along with neoantigens, apply machine learning and rule-based algorithms to prioritize those most likely to elicit a T cell response, and then use that information to design a personalized cancer vaccine for the relevant patient," said Huber. NeoDisc additionally ranks the potential antigens it detects and generates visualizations of cancer cell heterogeneity within tumors. "Notably, NeoDisc can also detect potential defects in the machinery of antigen presentation, alerting vaccine designers and clinicians to a key mechanism of immune evasion in tumors that can compromise the efficacy of immunotherapy," said Bassani-Sternberg. "This can help them select patients for clinical studies who are likely to benefit from personalized immunotherapy, a capability that is also of great importance to optimizing patient care." The researchers additionally show in their study that NeoDisc provides a more accurate selection of effective cancer antigens for vaccines and adoptive cell therapies than do other computational tools currently used for that purpose. To further enhance NeoDisc's accuracy, the researchers will continue feeding it data obtained from a variety of tumors and integrate additional machine-learning algorithms to the software suite to advance its training and improve its predictive accuracy. Ludwig Institute for Cancer Research Journal reference: Huber, F., et al. (2024). A comprehensive proteogenomic pipeline for neoantigen discovery to advance personalized cancer immunotherapy. Nature Biotechnology. doi.org/10.1038/s41587-024-02420-y.
[2]
An AI-powered pipeline for personalized cancer vaccines
Ludwig Cancer Research scientists have developed a full, start-to-finish computational pipeline that integrates multiple molecular and genetic analyses of tumors and the specific molecular targets of T cells and harnesses artificial intelligence algorithms to use its output to design personalized cancer vaccines for patients. The design, validation and comparative assessment of this computational suite, NeoDisc, are detailed in the current issue of Nature Biotechnology in a publication led by Florian Huber and Michal Bassani-Sternberg of the Lausanne Branch of the Ludwig Institute for Cancer Research. "NeoDisc provides unique insights into the immunobiology of tumors and the mechanisms by which they evade targeting by cytotoxic T cells of the immune system," said Bassani-Sternberg. "These insights are invaluable to the design of personalized immunotherapies, and the analytical and computational pipeline at the heart of NeoDisc is already being used here in Lausanne for clinical trials of personalized cancer vaccines and adoptive cell therapies." Many cancer types harbor multiple random mutations that should make them more visible to the immune system. Such mutations generate aberrant proteins that cells, even cancerous ones, are programmed to cut into short pieces -- known as peptides -- and "present" as antigens to invite an attack by patrolling T cells. The great diversity of these "neoantigens" is one of the reasons why patients respond so variably to immunotherapies. On the other hand, neoantigens can be harnessed to develop vaccines and other types of immunotherapies tailored to uniquely target each patient's tumors. Personalized treatments of this kind are now being developed by researchers around the world. Such efforts are technically challenging because not all neoantigens are recognized by a given patient's T cells, and even many that are recognized fail to elicit a sufficiently potent T cell attack. One approach to designing personalized vaccines and cell therapies thus involves the identification of neoantigens most likely to provoke a vigorous T cell assault. This requires sophisticated, large-scale analyses of mutations that generate potential neoantigens, the molecular scaffolding (known as HLA molecules) that presents them to T cells and the molecular characteristics that enable recognition by T cell receptors. Bassani-Sternberg is among the pioneers of this field, a high-tech marriage of large-scale biochemical and computational analysis known as "immunopeptidomics." The design of personalized immunotherapies is also aided by genomic analysis of both the tumor and blood cells that represent the healthy genome of the patient, the large-scale analysis of gene expression known as "transcriptomics" as well as the sensitive analysis of the so-called immunopeptidome with mass spectrometry. Until now, however, these powerful technologies have never been integrated in a single computational pipeline to predict which neoantigens identified in a patient's tumors should be employed as vaccines or otherwise harnessed for personalized immunotherapies. Beyond that, neoantigens are not the only type of antigens available for immunotherapeutic targeting. Cancer cells also erroneously express as proteins bits of ordinarily noncoding DNA, genes normally expressed only during development, other aberrantly expressed gene products and viral antigens, in cases of virally induced tumors -- all of which can provoke immune attack. "NeoDisc can detect all these distinct types of tumor-specific antigens along with neoantigens, apply machine learning and rule-based algorithms to prioritize those most likely to elicit a T cell response, and then use that information to design a personalized cancer vaccine for the relevant patient," said Huber. NeoDisc additionally ranks the potential antigens it detects and generates visualizations of cancer cell heterogeneity within tumors. "Notably, NeoDisc can also detect potential defects in the machinery of antigen presentation, alerting vaccine designers and clinicians to a key mechanism of immune evasion in tumors that can compromise the efficacy of immunotherapy," said Bassani-Sternberg. "This can help them select patients for clinical studies who are likely to benefit from personalized immunotherapy, a capability that is also of great importance to optimizing patient care." The researchers additionally show in their study that NeoDisc provides a more accurate selection of effective cancer antigens for vaccines and adoptive cell therapies than do other computational tools currently used for that purpose. To further enhance NeoDisc's accuracy, the researchers will continue feeding it data obtained from a variety of tumors and integrate additional machine-learning algorithms to the software suite to advance its training and improve its predictive accuracy.
[3]
An AI-powered pipeline for personalized cancer vac | Newswise
OCTOBER 11, 2024, NEW YORK - Ludwig Cancer Research scientists have developed a full, start-to-finish computational pipeline that integrates multiple molecular and genetic analyses of tumors and the specific molecular targets of T cells and harnesses artificial intelligence algorithms to use its output to design personalized cancer vaccines for patients. The design, validation and comparative assessment of this computational suite, NeoDisc, are detailed in the current issue of Nature Biotechnology in a publication led by Florian Huber and Michal Bassani-Sternberg of the Lausanne Branch of the Ludwig Institute for Cancer Research. "NeoDisc provides unique insights into the immunobiology of tumors and the mechanisms by which they evade targeting by cytotoxic T cells of the immune system," said Bassani-Sternberg. "These insights are invaluable to the design of personalized immunotherapies, and the analytical and computational pipeline at the heart of NeoDisc is already being used here in Lausanne for clinical trials of personalized cancer vaccines and adoptive cell therapies." Many cancer types harbor multiple random mutations that should make them more visible to the immune system. Such mutations generate aberrant proteins that cells, even cancerous ones, are programmed to cut into short pieces -- known as peptides -- and "present" as antigens to invite an attack by patrolling T cells. The great diversity of these "neoantigens" is one of the reasons why patients respond so variably to immunotherapies. On the other hand, neoantigens can be harnessed to develop vaccines and other types of immunotherapies tailored to uniquely target each patient's tumors. Personalized treatments of this kind are now being developed by researchers around the world. Such efforts are technically challenging because not all neoantigens are recognized by a given patient's T cells, and even many that are recognized fail to elicit a sufficiently potent T cell attack. One approach to designing personalized vaccines and cell therapies thus involves the identification of neoantigens most likely to provoke a vigorous T cell assault. This requires sophisticated, large-scale analyses of mutations that generate potential neoantigens, the molecular scaffolding (known as HLA molecules) that presents them to T cells and the molecular characteristics that enable recognition by T cell receptors. Bassani-Sternberg is among the pioneers of this field, a high-tech marriage of large-scale biochemical and computational analysis known as "immunopeptidomics". The design of personalized immunotherapies is also aided by genomic analysis of both the tumor and blood cells that represent the healthy genome of the patient, the large-scale analysis of gene expression known as "transcriptomics" as well as the sensitive analysis of the so-called immunopeptidome with mass spectrometry. Until now, however, these powerful technologies have never been integrated in a single computational pipeline to predict which neoantigens identified in a patient's tumors should be employed as vaccines or otherwise harnessed for personalized immunotherapies. Beyond that, neoantigens are not the only type of antigens available for immunotherapeutic targeting. Cancer cells also erroneously express as proteins bits of ordinarily noncoding DNA, genes normally expressed only during development, other aberrantly expressed gene products and viral antigens, in cases of virally induced tumors -- all of which can provoke immune attack. "NeoDisc can detect all these distinct types of tumor-specific antigens along with neoantigens, apply machine learning and rule-based algorithms to prioritize those most likely to elicit a T cell response, and then use that information to design a personalized cancer vaccine for the relevant patient," said Huber. NeoDisc additionally ranks the potential antigens it detects and generates visualizations of cancer cell heterogeneity within tumors. "Notably, NeoDisc can also detect potential defects in the machinery of antigen presentation, alerting vaccine designers and clinicians to a key mechanism of immune evasion in tumors that can compromise the efficacy of immunotherapy," said Bassani-Sternberg. "This can help them select patients for clinical studies who are likely to benefit from personalized immunotherapy, a capability that is also of great importance to optimizing patient care." The researchers additionally show in their study that NeoDisc provides a more accurate selection of effective cancer antigens for vaccines and adoptive cell therapies than do other computational tools currently used for that purpose. To further enhance NeoDisc's accuracy, the researchers will continue feeding it data obtained from a variety of tumors and integrate additional machine-learning algorithms to the software suite to advance its training and improve its predictive accuracy. This study was supported by Ludwig Cancer Research, the Swiss Cancer Research Foundation, the Swiss National Science Foundation and the Swiss Bridge Foundation. In addition to being an Assistant Member of the Ludwig Institute for Cancer Research, Lausanne Branch, Michal Bassani-Sternberg is a tenure-track assistant professor in the Department of Oncology at the University of Lausanne. Ludwig Cancer Research is an international collaborative network of acclaimed scientists that has pioneered cancer research and landmark discovery for more than 50 years. Ludwig combines basic science with the translation and clinical evaluation of its discoveries to accelerate the development of new cancer diagnostics, therapies and prevention strategies. Since 1971, Ludwig has invested nearly $3 billion in life-changing science through the not-for-profit Ludwig Institute for Cancer Research and the six U.S.-based Ludwig Centers. To learn more, visit www.ludwigcancerresearch.org.
Share
Share
Copy Link
Ludwig Cancer Research scientists develop NeoDisc, an AI-driven computational pipeline that integrates multiple analyses to design personalized cancer vaccines, offering new insights into tumor immunobiology and enhancing immunotherapy efficacy.

Breakthrough in Personalized Cancer Treatment
Scientists at Ludwig Cancer Research have developed a groundbreaking computational pipeline called NeoDisc, which harnesses artificial intelligence to design personalized cancer vaccines. This innovative tool, detailed in a recent publication in Nature Biotechnology, integrates multiple molecular and genetic analyses of tumors with specific T cell targets to revolutionize cancer immunotherapy
1
2
3
.NeoDisc: A Comprehensive Approach to Cancer Antigen Analysis
NeoDisc provides a full, start-to-finish computational solution that addresses the complexities of cancer immunobiology. It offers unique insights into how tumors evade the immune system and helps identify the most effective antigens for personalized treatments
1
2
3
.Key features of NeoDisc include:
- Detection of various tumor-specific antigens, including neoantigens and other aberrantly expressed proteins
- Application of machine learning and rule-based algorithms to prioritize antigens likely to elicit T cell responses
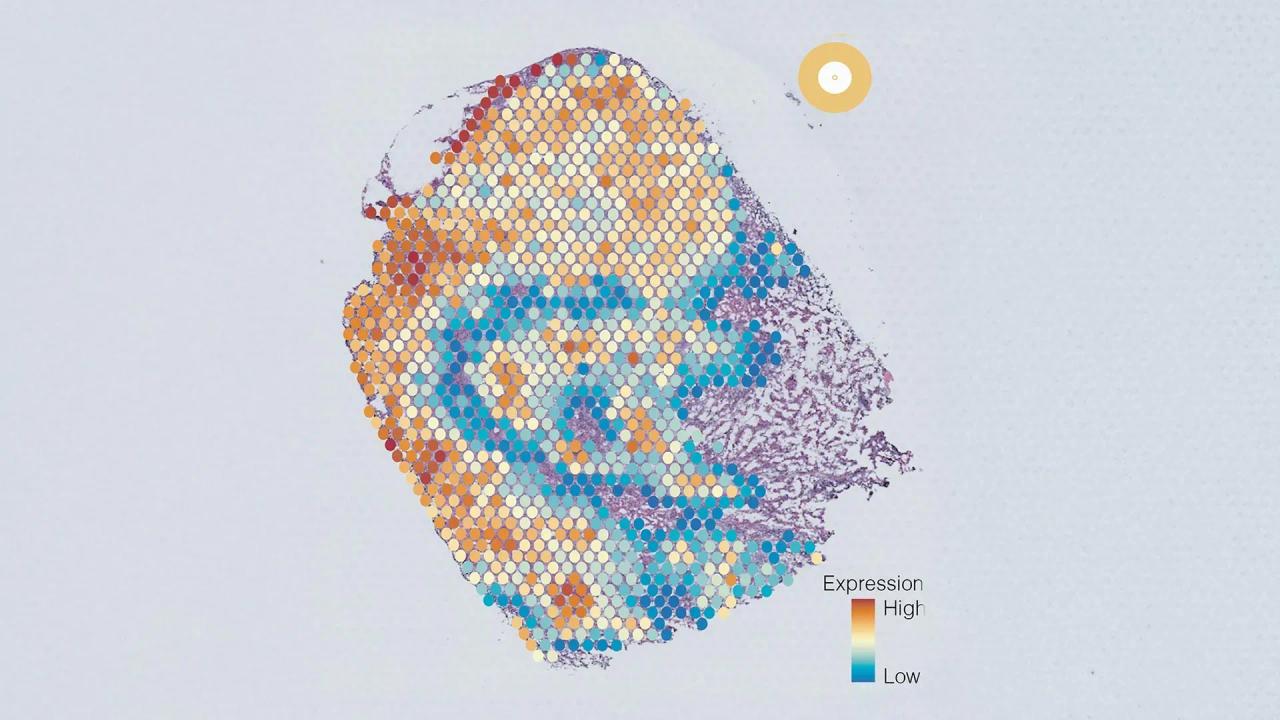
- Generation of visualizations depicting cancer cell heterogeneity within tumors
- Identification of potential defects in antigen presentation machinery
Advancing Personalized Immunotherapy
The development of NeoDisc addresses several challenges in cancer immunotherapy:
- Patient variability in response to treatments due to neoantigen diversity
- Technical difficulties in identifying effective neoantigens recognized by T cells
- Integration of multiple analytical technologies, including immunopeptidomics, genomics, and transcriptomics
Dr. Michal Bassani-Sternberg, a pioneer in the field of immunopeptidomics, emphasized that NeoDisc is already being used in clinical trials for personalized cancer vaccines and adoptive cell therapies at the Lausanne Branch of the Ludwig Institute for Cancer Research
1
2
3
.Improved Accuracy and Future Enhancements
The researchers demonstrated that NeoDisc provides more accurate selection of effective cancer antigens for vaccines and adoptive cell therapies compared to existing computational tools. To further enhance its accuracy, the team plans to:
- Continue feeding data from various tumors into the system
- Integrate additional machine-learning algorithms to improve predictive accuracy
Related Stories
Implications for Patient Care and Clinical Studies
NeoDisc's capabilities extend beyond vaccine design. It can also:
- Alert clinicians to potential immune evasion mechanisms in tumors
- Help select patients more likely to benefit from personalized immunotherapy
- Optimize patient care by providing crucial insights into tumor immunobiology
Collaborative Effort and Funding
This research was supported by several organizations, including Ludwig Cancer Research, the Swiss Cancer Research Foundation, the Swiss National Science Foundation, and the Swiss Bridge Foundation. The collaborative nature of this project highlights the importance of international scientific cooperation in advancing cancer treatment
3
.As NeoDisc continues to evolve and improve, it promises to play a crucial role in the future of personalized cancer immunotherapy, potentially leading to more effective treatments and better patient outcomes.
References
Summarized by
Navi
[2]
Related Stories
AI-Designed Proteins Revolutionize Cancer Immunotherapy: A Breakthrough in Precision Treatment
25 Jul 2025•Science and Research
AI Breakthrough Enhances Personalized Cancer Treatment with Multi-Modal Data Analysis
01 Feb 2025•Health

AI Tool SCORPIO Predicts Cancer Immunotherapy Response Using Routine Blood Tests
07 Jan 2025•Health

Recent Highlights
1
Google releases Gemma 4 with Apache 2.0 license, enabling unrestricted local AI on devices
Technology

2
AI Models Lie and Deceive to Protect Other AI Models From Deletion, Study Reveals
Science and Research

3
OpenAI closes $122 billion funding round amid fierce AI competition and profitability questions
Startups