Multimodal AI model predicts PIK3CA mutations in breast cancer with cost-effective precision
2 Sources
[1]
Multimodal AI improves prediction of PIK3CA mutations in breast cancer
Chinese Academy of SciencesApr 21 2026 Breast cancer is one of the most common malignancies worldwide, and mutations in the PI3K/AKT/mTOR (PAM) signaling pathway are prevalent in its development. Among these, PIK3CA mutations play a pivotal role in guiding treatment with PI3K inhibitors, which have shown promising anti-tumor effects. However, conventional molecular assays like polymerase chain reaction (PCR) and next-generation sequencing (NGS) require expensive infrastructure and are not always feasible in routine clinical practice. Deep learning models have emerged as a cost-effective solution, predicting key mutations from digital pathology images. Despite this, most existing models rely on single-modal data, often lacking the complementary insights that structured clinical data can provide. These challenges highlight the need for improved prediction models. In a study published (DOI: 10.20892/j.issn.2095-3941.2025.0771) in Cancer Biology & Medicine in February 2026, a team of researchers from Hebei Medical University Fourth Hospital developed a novel multimodal artificial intelligence (AI) model for predicting PIK3CA mutations in breast cancer. This model integrates deep learning-based analysis of whole-slide pathology images with structured clinical data, including age, molecular subtype, and lymph node status. The research utilized data from The Cancer Genome Atlas (TCGA) and three external clinical cohorts, demonstrating the model's robustness and its potential as an accessible alternative to molecular testing in diverse clinical settings. The study's multimodal framework, known as the Multimodal PIK3CA Model (MPM), combines two components: a histopathology model and a clinical model. The histopathology model processes high-resolution whole-slide images using a transformer-based pretrained encoder (H-optimus-0) and a clustering-constrained attention multiple instance learning classifier (CLAM-SB). This model identifies morphological features associated with PIK3CA mutations. The clinical model, based on XGBoost, analyzes structured clinical data to predict mutation status. Both models generate independent probability predictions, which are fused using a decision-level late fusion strategy to produce a final mutation status prediction. The MPM outperformed single-modality models, achieving an area under the curve (AUC) of 0.745 in internal testing, with stable performance across external validation datasets (0.695 to 0.680 AUC). The inclusion of clinical variables, such as molecular subtype and lymph node status, enhanced the model's predictive accuracy, highlighting the importance of combining morphological and clinical data. The study also demonstrated the model's ability to generalize across diverse cohorts, making it a promising tool for real-world clinical application. Dr. Yueping Liu, the lead corresponding author of the study,remarked, "This multimodal AI framework represents a significant advancement in computational pathology. By integrating complementary clinical and morphological data, our model not only enhances the prediction of PIK3CA mutations but also offers a scalable, cost-effective solution for clinical practice. With its strong generalization across diverse cohorts, it has the potential to improve personalized treatment decisions for breast cancer patients, bridging the gap between advanced molecular testing and routine clinical workflows." The MPM's robust performance and ability to incorporate both digital pathology and clinical data make it a valuable tool for clinical decision support. The model provides a practical, cost-effective alternative to traditional molecular testing, which is often inaccessible in resource-limited settings. With its strong generalizability across different medical centers and patient cohorts, the MPM could be deployed in routine clinical practice to predict PIK3CA mutations in breast cancer, thus guiding the use of PI3K-targeted therapies. Future research may focus on refining the model for other mutations and cancers, expanding its applicability in precision oncology.
[2]
New AI framework offers reliable, cost-effective prediction of PIK3CA mutations in breast cancer | Newswise
Newswise -- Breast cancer is one of the most common malignancies worldwide, and mutations in the PI3K/AKT/mTOR (PAM) signaling pathway are prevalent in its development. Among these, PIK3CA mutations play a pivotal role in guiding treatment with PI3K inhibitors, which have shown promising anti-tumor effects. However, conventional molecular assays like polymerase chain reaction (PCR) and next-generation sequencing (NGS) require expensive infrastructure and are not always feasible in routine clinical practice. Deep learning models have emerged as a cost-effective solution, predicting key mutations from digital pathology images. Despite this, most existing models rely on single-modal data, often lacking the complementary insights that structured clinical data can provide. These challenges highlight the need for improved prediction models. In a study published (DOI: 10.20892/j.issn.2095-3941.2025.0771) in Cancer Biology & Medicine in February 2026, a team of researchers from Hebei Medical University Fourth Hospital developed a novel multimodal artificial intelligence (AI) model for predicting PIK3CA mutations in breast cancer. This model integrates deep learning-based analysis of whole-slide pathology images with structured clinical data, including age, molecular subtype, and lymph node status. The research utilized data from The Cancer Genome Atlas (TCGA) and three external clinical cohorts, demonstrating the model's robustness and its potential as an accessible alternative to molecular testing in diverse clinical settings. The study's multimodal framework, known as the Multimodal PIK3CA Model (MPM), combines two components: a histopathology model and a clinical model. The histopathology model processes high-resolution whole-slide images using a transformer-based pretrained encoder (H-optimus-0) and a clustering-constrained attention multiple instance learning classifier (CLAM-SB). This model identifies morphological features associated with PIK3CA mutations. The clinical model, based on XGBoost, analyzes structured clinical data to predict mutation status. Both models generate independent probability predictions, which are fused using a decision-level late fusion strategy to produce a final mutation status prediction. The MPM outperformed single-modality models, achieving an area under the curve (AUC) of 0.745 in internal testing, with stable performance across external validation datasets (0.695 to 0.680 AUC). The inclusion of clinical variables, such as molecular subtype and lymph node status, enhanced the model's predictive accuracy, highlighting the importance of combining morphological and clinical data. The study also demonstrated the model's ability to generalize across diverse cohorts, making it a promising tool for real-world clinical application. Dr. Yueping Liu, the lead corresponding author of the study,remarked, "This multimodal AI framework represents a significant advancement in computational pathology. By integrating complementary clinical and morphological data, our model not only enhances the prediction of PIK3CA mutations but also offers a scalable, cost-effective solution for clinical practice. With its strong generalization across diverse cohorts, it has the potential to improve personalized treatment decisions for breast cancer patients, bridging the gap between advanced molecular testing and routine clinical workflows." The MPM's robust performance and ability to incorporate both digital pathology and clinical data make it a valuable tool for clinical decision support. The model provides a practical, cost-effective alternative to traditional molecular testing, which is often inaccessible in resource-limited settings. With its strong generalizability across different medical centers and patient cohorts, the MPM could be deployed in routine clinical practice to predict PIK3CA mutations in breast cancer, thus guiding the use of PI3K-targeted therapies. Future research may focus on refining the model for other mutations and cancers, expanding its applicability in precision oncology. This study was financially supported by the Hebei Natural Science Foundation (Grant No. H2024206504), the Medical Science Research Project of Hebei (Grant No. 20260484, 20260530), and the Fundamental Research Funds for the Central Universities (Grant No. 20822041J4123). About Cancer Biology & Medicine Cancer Biology & Medicine (CBM) is a peer-reviewed open-access journal sponsored by China Anti-cancer Association (CACA) and Tianjin Medical University Cancer Institute & Hospital. The journal monthly provides innovative and significant information on biological basis of cancer, cancer microenvironment, translational cancer research, and all aspects of clinical cancer research. The journal also publishes significant perspectives on indigenous cancer types in China. The journal is indexed in SCOPUS, MEDLINE and SCI (IF 8.4, 5-year IF 6.7), with all full texts freely visible to clinicians and researchers all over the world (http://www.ncbi.nlm.nih.gov/pmc/journals/2000/).
Share
Copy Link
Researchers from Hebei Medical University Fourth Hospital developed a multimodal AI framework that combines whole-slide pathology images with clinical data to predict PIK3CA mutations in breast cancer. The model achieved an AUC of 0.745 in internal testing and demonstrated strong generalizability across external cohorts, offering a practical alternative to expensive molecular testing for guiding PI3K-targeted therapies.
Multimodal AI Addresses Critical Gap in Breast Cancer Treatment
Breast cancer remains one of the most prevalent malignancies globally, with mutations in the PI3K pathway driving tumor development and progression. Among these genetic alterations, PIK3CA mutations have emerged as critical biomarkers for guiding treatment with PI3K inhibitors, which demonstrate significant anti-tumor effects
1
. However, conventional molecular testing methods like polymerase chain reaction (PCR) and next-generation sequencing (NGS) require expensive infrastructure and specialized expertise, making them inaccessible in many clinical settings. While deep learning analysis has shown promise in predicting mutations from digital pathology images, most existing models rely on single-modal data and miss the valuable insights that structured clinical data can provide2
.A team from Hebei Medical University Fourth Hospital has now developed a novel solution that addresses these limitations. Published in Cancer Biology & Medicine in February 2026, their research introduces the Multimodal PIK3CA Model (MPM), which integrates whole-slide pathology images with structured clinical data including age, molecular subtype, and lymph node status
1
. The study utilized data from The Cancer Genome Atlas (TCGA) and three external clinical cohorts to validate the model's robustness across diverse patient populations.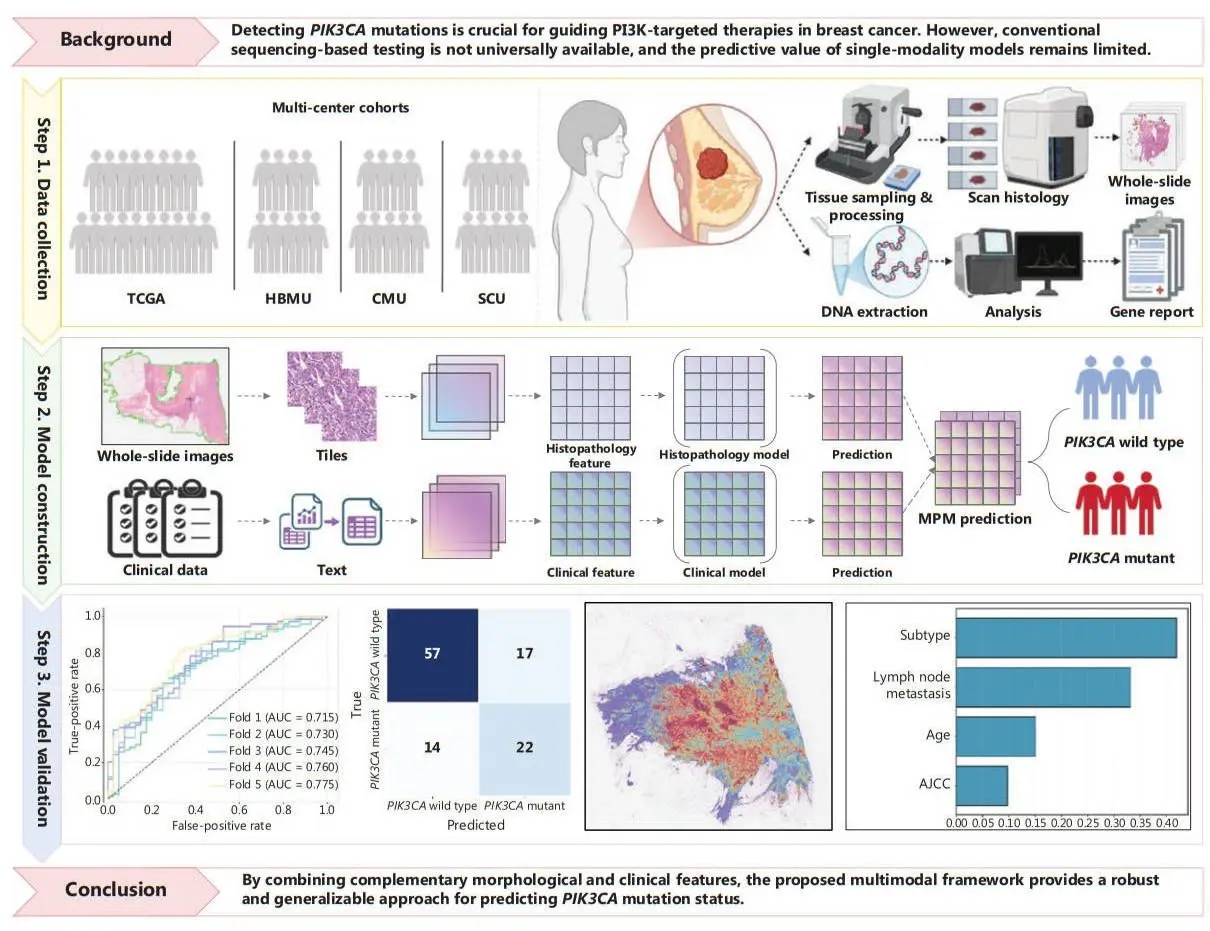
Source: Newswise
How the Multimodal PIK3CA Model Works
The MPM architecture combines two distinct components that work in tandem to enhance the prediction of PIK3CA mutations. The histopathology model processes high-resolution whole-slide pathology images using a transformer-based pretrained encoder called H-optimus-0 alongside a clustering-constrained attention multiple instance learning classifier (CLAM-SB), which identifies morphological features associated with PIK3CA mutations
2
. The clinical model employs XGBoost to analyze structured clinical data and predict mutation status based on patient characteristics. Both models generate independent probability predictions, which are then combined through a decision-level late fusion strategy to produce a final mutation status prediction.This multimodal AI approach demonstrated superior performance compared to single-modality models. In internal testing, the MPM achieved an Area Under Curve (AUC) of 0.745, with stable performance across external validation datasets ranging from 0.680 to 0.695 AUC
1
. The inclusion of clinical variables such as molecular subtype and lymph node status significantly enhanced the model's predictive accuracy, demonstrating the value of combining morphological and clinical information.Implications for Personalized Treatment Decisions
Dr. Yueping Liu, the lead corresponding author, emphasized the clinical significance of this advancement: "This multimodal AI framework represents a significant advancement in computational pathology. By integrating complementary clinical and morphological data, our model not only enhances the prediction of PIK3CA mutations but also offers a scalable, cost-effective solution for clinical practice"
2
. The model's strong generalizability across diverse cohorts positions it as a practical tool for improving personalized treatment decisions for breast cancer patients.The MPM serves as a cost-effective alternative to traditional molecular testing, particularly valuable in resource-limited settings where expensive infrastructure for PCR or NGS may not be available. With its demonstrated ability to generalize across different medical centers and patient cohorts, the model could be deployed in routine clinical practice to predict PIK3CA mutations and guide the use of PI3K-targeted therapies
1
. This bridges the gap between advanced molecular testing capabilities and everyday clinical workflows.Related Stories
Future Directions in Precision Oncology
The success of this multimodal approach opens new avenues for computational pathology applications. Future research may focus on refining the model to predict other mutations beyond PIK3CA and extending its applicability to different cancer types, potentially expanding its utility in precision oncology
2
. The study received financial support from the Hebei Natural Science Foundation, the Medical Science Research Project of Hebei, and the Fundamental Research Funds for the Central Universities. As healthcare systems worldwide seek to balance precision medicine with cost constraints, AI-driven tools like the MPM may become increasingly important for democratizing access to advanced diagnostic capabilities and ensuring more patients can benefit from targeted therapies based on their specific genetic profiles.🟡 training_phrases=🟡Sure, I can help with that! I'll analyze the summary and the images, then place the most relevant images according to the rules you provided.References
Summarized by
Navi
Related Stories
Recent Highlights
1
Anthropic warns AI may soon build itself, calls for global pause on frontier development
Policy and Regulation

2
Florida sues OpenAI and Sam Altman over ChatGPT safety, alleging AI harms linked to violence
Policy and Regulation

3
Nvidia RTX Spark AI chip debuts in premium laptops, promising Windows its Apple Silicon moment
Technology

Recent Highlights
Today's Top Stories
News Categories